- HOME
- Laboratories
- Molecular Pharmacology
- Projects
Projects
- Development of "Cancer Cell Informatics", an information-rich platform for the research of molecular-targeted therapy
- Discovery and development of novel molecular-targeted drugs
- Development of PI3 kinase-targeted drugs
- Molecular pharmacology of anticancer drugs based on chemical and genomic information
Development of "Cancer Cell Informatics", an information-rich platform for the research of molecular-targeted therapy
We established a panel of human cancer cell lines coupled to a database integrating their chemosensitivities and genomic information, termed “Cancer Cell Informatics (CCI)”, which is an information-rich system for studying molecular-targeted therapy. It is most powerful in drug discovery and in predicting molecular targets of new chemicals. Furthermore, it potentially provides valuable information for studying molecular pharmacology of new drugs and for identifying various biomarkers and new targets.
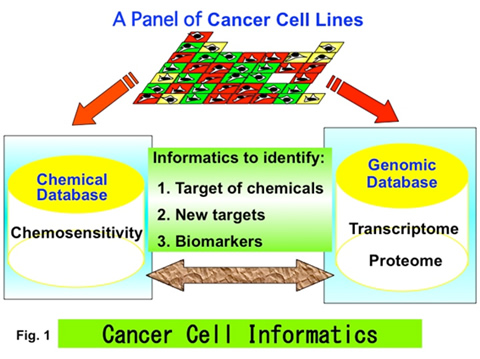
Fig.1 Cancer Cell Informatics(CCI)
Cancer Cell Informatics is a system based on a database integrating the chemosensitivities, gene- and protein expression profiles of a panel of 39 human cancer cell lines (JFCR39), which provides information on the relation between chemicals and gene products. For example, it is powerful in predicting a potential molecular target of a chemical of interest.
Discovery and development of novel molecular-targeted drugs
Discovering new anticancer compounds is important to develop anticancer drugs. We are conducting anticancer drug screening based on CCI described above. We evaluate test compounds based on their growth-inhibitory activity across a panel of 39 human cancer cell lines (termed JFCR39) using a data-mining tool “COMPARE”, which is comparable to NCI-60 developed by National Cancer Institute in USA and potentially predicts the molecular targets of compounds. We indeed identified novel anticancer compounds, for example, MS-247 (an inhibitor of topoisomerase I / II), pladienolide B (a lead of inhibitors of splicing factor SF3B) and ZSTK474 (an inhibitor of PI3 kinase).
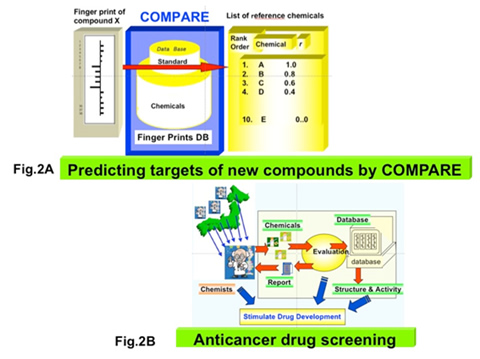
Fig.2: Prediction of the molecular target of a chemical and screening of anticancer drugs
A) Once the activity profile of compound X across JFCR39 (termed Fingerprint) is measured, the molecular target or the mode of action of X can be predicted by COMPARE algorithm that verifies the similarity of X to a certain reference compound in terms of Fingerprints. B) In collaboration with other institutes, we screen the compounds with unique molecular targets.
Development of PI3 kinase-targeted drugs
PI3K (Phosphatidylinositol 3-kinase) is a key enzyme in signal transduction and plays important roles in survival, growth and metastasis of cancer. Therefore, PI3K is a promising target for cancer therapy. We identified ZSTK474, a novel PI3K inhibitor, which has potent anticancer activity via oral administration without severe toxicity. We are doing the translational research of ZSTK474 to indicate its potential efficacy in various cancer models and to identify the biomarkers for predicting its efficacy.
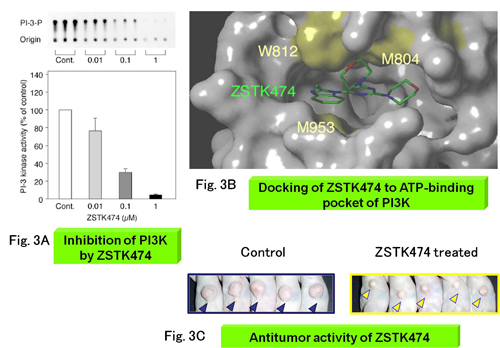
Fig.3 Identification of a new PI3K inhibitor ZSTK474
A) We predicted that the target of a novel compound ZSTK474 might be PI3K by CCI, and experimentally demonstrated that the prediction was true. B) We demonstrated that ZSTK474 binds to the ATP-binding pocket of PI3K by molecular modeling and biological experiments. C) ZSTK474 showed potent therapeutic efficacy against human cancer xenografts.
Molecular pharmacology of anticancer drugs based on chemical and genomic information
We conduct identification of new molecular targets for anticancer therapy by using CCI. We also try to identify the genes determining chemosensitivity in cancer using RNAi-based gene knockdown method.
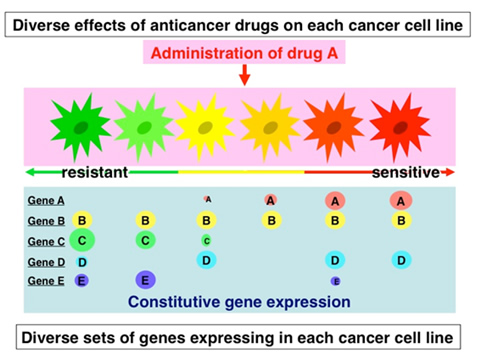
Fig.4 Identification of the genes determining chemosensitivity in cancer
We try to identify the genes determining chemosensitivity in cancer by mining the data of chemosensitivity and gene expression profiles. If Gene A, for example, is highly expressed in highly Drug X-sensitive cancers, Gene A may be involved in the chemosensitivity of cancer to Drug X. On the contrary, if Gene C is highly expressed in highly Drug X-resistant cancers, Gene C may be involved in the resistance of cancer to Drug X.





